(UPDATED: See below.)
 As I discussed a few weeks ago with respect to deCODEme — a “personal genomics” service hurriedly launched last November by Iceland’s deCODE Genetics in an apparent attempt to beat 23andMe to market (it succeeded by a day or so) — these sorts of services can awfully dense and difficult to navigate. The deCODEme service appears to be particularly bad in that respect, both in terms of its design and even the underlying science used to justify the genetic information displayed in a demonstration user account.
As I discussed a few weeks ago with respect to deCODEme — a “personal genomics” service hurriedly launched last November by Iceland’s deCODE Genetics in an apparent attempt to beat 23andMe to market (it succeeded by a day or so) — these sorts of services can awfully dense and difficult to navigate. The deCODEme service appears to be particularly bad in that respect, both in terms of its design and even the underlying science used to justify the genetic information displayed in a demonstration user account.
So it’s a pleasant surprise to report that 23andMe, which over the weekend began allowing people to set up demonstration accounts itself, appears to have made the process of understanding your genetic inheritance about as simple and intuitive as it can probably get. The demo accounts don’t display your own genetic information, of course — instead, they show a profile for the fictional Greg and Lilly Mendel and their immediate relatives. (The family name is inspired by Gregor Mendel, a nineteenth-century monk known as the “father of genetics” for his studies on the inheritance of pea plants; the profile uses actual data from an anonymous European family.)
These sorts of demo accounts are particularly useful given that 23andMe and its competitors are charging customers roughly $1,000 for a genetic analysis, which is a lot to shell out when you don’t have any real idea what you’re getting for your money. To sign up for a 23andMe demo account, click here.
AI Weekly
The must-read newsletter for AI and Big Data industry written by Khari Johnson, Kyle Wiggers, and Seth Colaner.
Included with VentureBeat Insider and VentureBeat VIP memberships.
 Like all these services — of which there are currently at least four, counting the new DNATraits project launched recently by Family Tree DNA — 23andMe takes a genetic sample (here from having users spit repeatedly into a tube) and checks 600,000 or so individual DNA “letters,” or bases, known to vary between people. After analyzing those letters, the company posts your genetic information on a Web site where you can see what your particular genetic pattern says about inherited traits such as your susceptibility to cancer or heart disease, longevity and even eye and hair color. Not only can you spin through the data in as much or as little detail as you like, you can share it with relatives or friends and search for others with similar traits.
Like all these services — of which there are currently at least four, counting the new DNATraits project launched recently by Family Tree DNA — 23andMe takes a genetic sample (here from having users spit repeatedly into a tube) and checks 600,000 or so individual DNA “letters,” or bases, known to vary between people. After analyzing those letters, the company posts your genetic information on a Web site where you can see what your particular genetic pattern says about inherited traits such as your susceptibility to cancer or heart disease, longevity and even eye and hair color. Not only can you spin through the data in as much or as little detail as you like, you can share it with relatives or friends and search for others with similar traits.
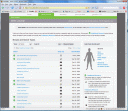
One of the first things a new visitor will see is the clean and uncluttered look of 23andMe’s “gene journal,” which lets you scroll through various genetic traits and then dive in to see how you — well, your Mendel stand-in — fare compared to the population at large. (See a screenshot of Greg Mendel’s gene journal using the thumbnail above and to the left.)
More after the jump:
 The contrast with deCODEme couldn’t be more striking. Where the Icelandic service offers minimal data and requires lots of clicking just to see how your genes stack up against those of the general population, 23andMe puts almost all that information out front. Each condition is rated according to the reliability of the data that links particular genetic variations (technically known as “single nucleotide polymorphisms,” or SNPs) — I’ve blown up the ratings in the graphic to the left; click for a larger version — which itself is a big step, since deCODEme appears to limit what it shows customers to the most conservative data available. Yes, it’s always possible that people will misinterpret their disease risk based on sketchy data, but here such mistakes are particularly difficult to make, given that the site deploys simple icons and contant boxed reminders whenever gene-disease links are considered preliminary as opposed to established.
The contrast with deCODEme couldn’t be more striking. Where the Icelandic service offers minimal data and requires lots of clicking just to see how your genes stack up against those of the general population, 23andMe puts almost all that information out front. Each condition is rated according to the reliability of the data that links particular genetic variations (technically known as “single nucleotide polymorphisms,” or SNPs) — I’ve blown up the ratings in the graphic to the left; click for a larger version — which itself is a big step, since deCODEme appears to limit what it shows customers to the most conservative data available. Yes, it’s always possible that people will misinterpret their disease risk based on sketchy data, but here such mistakes are particularly difficult to make, given that the site deploys simple icons and contant boxed reminders whenever gene-disease links are considered preliminary as opposed to established.
You can also sort disease conditions by the affected region of the body — a feature that actually works here, as opposed to a similar feature at deCODEme when I last beamed in.
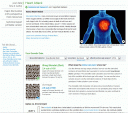 The individual disease-information pages open with some general information about the condition, but quickly get to the good stuff (here, I’ve chosen heart attack; click the thumbnail at left for a larger image). The site quickly presents you with your general risk versus that of a comparable population — in this example, European males aged 45-54, although you can adjust those settings via drop-down menus — then outlines the extent to which genes, as opposed to environment, are thought to contribute to the danger of heart attack.
The individual disease-information pages open with some general information about the condition, but quickly get to the good stuff (here, I’ve chosen heart attack; click the thumbnail at left for a larger image). The site quickly presents you with your general risk versus that of a comparable population — in this example, European males aged 45-54, although you can adjust those settings via drop-down menus — then outlines the extent to which genes, as opposed to environment, are thought to contribute to the danger of heart attack.
 Next — and this struck me as particularly cool — you get a graph of the individual SNPs 23andMe is using to make its calculations, which shows the positive and negative effects of each. Clicking on any particular SNP loads a panel immediately below that describes what’s known about it and lists citations from the scientific literature for more background. I could have sworn that the individual SNPs were also linked to a 23andMe database, allowing you to browse more deeply if you care to, but at the moment, if the disease pages permit that, I can’t find it again. You can click on a “technical report” link to the left in order to see the actual SNPs on which 23andMe bases its finding, but those aren’t linked either.
Next — and this struck me as particularly cool — you get a graph of the individual SNPs 23andMe is using to make its calculations, which shows the positive and negative effects of each. Clicking on any particular SNP loads a panel immediately below that describes what’s known about it and lists citations from the scientific literature for more background. I could have sworn that the individual SNPs were also linked to a 23andMe database, allowing you to browse more deeply if you care to, but at the moment, if the disease pages permit that, I can’t find it again. You can click on a “technical report” link to the left in order to see the actual SNPs on which 23andMe bases its finding, but those aren’t linked either.
Fortnately, unlike deCODEme, the 23andMe demo accounts are fully functional. The site’s GenomeExplorer lets you browse through your raw data, either in a graphical format displayed by chromosome or by searching on particular genes or SNPs. The output of such searches doesn’t strike me as terribly useful in its current incarnation, but that will presumably improve as time goes by. Instead, though, you can download “your” genetic data for use in other programs designed to read that format (such as this one). Don’t count on reading it directly without special help, though; I now have a 14MB file of “Greg Mendel’s” data sitting on my hard drive that’s simply too large to open with any common text-editing program.
You can also compare your genetics with others who share their information, although from my perspective, this feature is somewhat disappointing. Comparing Greg Mendel to his daughter, for instance, tells me only that they’re 84.15 percent similar across their entire genomes (at least that portion measured by 23andMe), or that across genes related to a limited number of characteristics — muscle endurance and circadian rhythm, for instance — they’re between 85 percent and 89 percent similar. This is probably more fun if you’re comparing yourself to your friends, assuming you’re open to that sort of thing, but I’m still at a loss as to exactly how this sort of stuff can prove useful.
I’ve sidestepped the entire ancestry section for now, although I’ll try to check back later if I have more time to look at it.
The 23andMe service isn’t without flaws, obviously. In many conditions, it also seems to limit the number of SNPs it reports on, much the same as I found at deCODEme. Heart attack is a prime example, since 23andMe omits the very same SNP strongly correlated with coronary-artery disease — rs1333049 — that I criticized deCODEme for skipping. And as noted above, some of the genetic exploration and comparison tools could probably use work. Overall, though, the service as displayed here is a pretty impressive effort.
But don’t take my word for it: Feel free to check out the 23andMe demo account yourself.
UPDATE: Review completed and rewritten throughout.
UPDATE REDUX: For the technically inclined, physician-turned-DNA enthusiast Ann Turner offers a comparative review of 23andMe and deCODEme over at Eye on DNA. (For what it’s worth, Turner also commented on my earlier deCODEme coverage here.)
Separately, 23andMe product manager Brian Naughton wrote to clarify that while 23andMe doesn’t use the rs1333049 SNP, it does use another SNP in the same chromosomal region that correlates highly (80 to 90 percent in Europeans) with rs1333049.
FURTHER UPDATE: deCODEme replies in comments. I’ll let their missive stand for itself, so take a look and see what you think.
VentureBeat's mission is to be a digital town square for technical decision-makers to gain knowledge about transformative enterprise technology and transact. Learn More